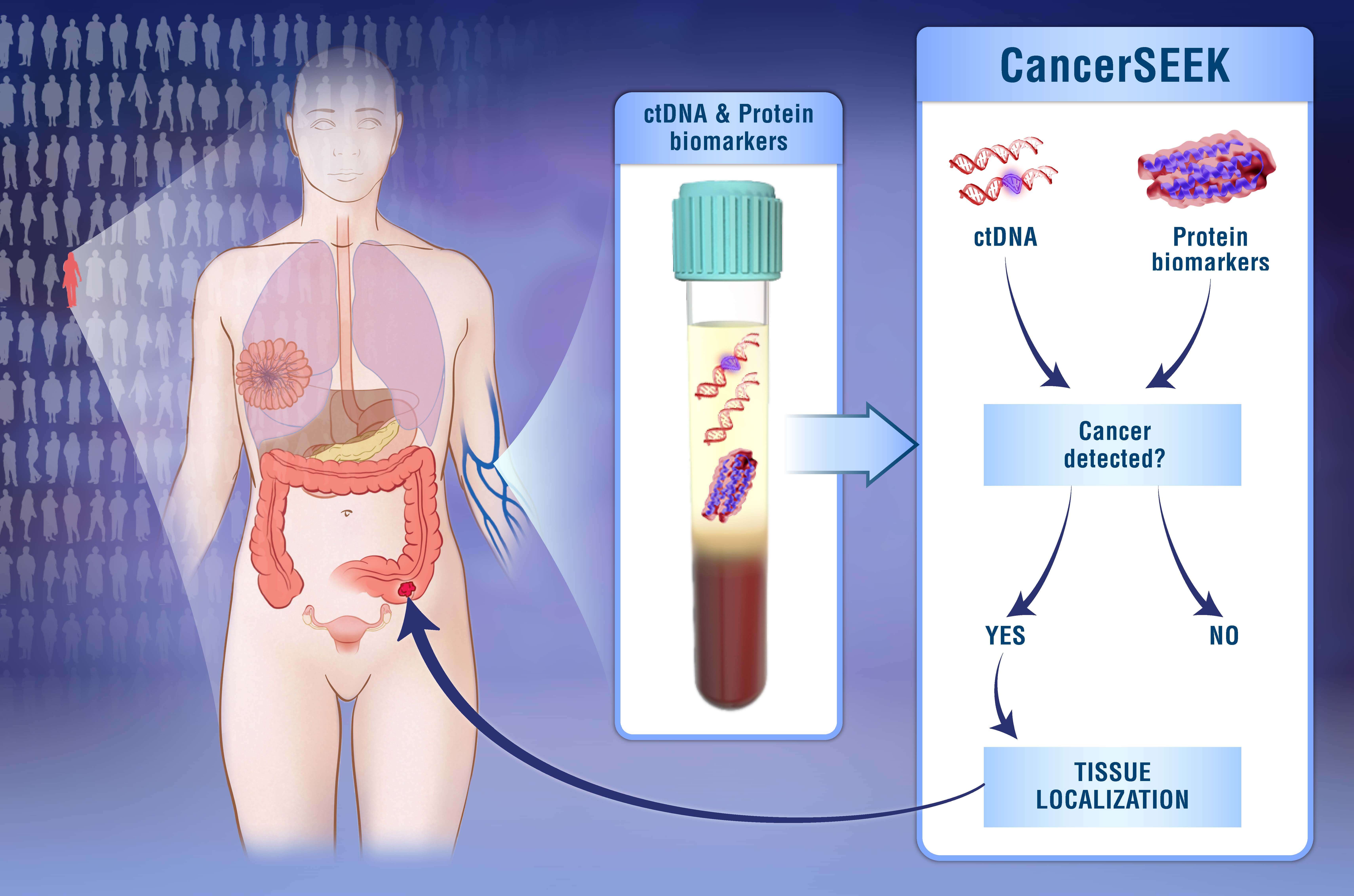
Credit: Elizabeth Cook and Kaitlin Lindsay
Johns Hopkins Kimmel Cancer Center researchers developed a single blood test that screens for eight common cancer types and helps identify the location of the cancer.
The test, called CancerSEEK, is a unique noninvasive, multianalyte test that simultaneously evaluates levels of eight cancer proteins and the presence of cancer gene mutations from circulating DNA in the blood. The test is aimed at screening for eight common cancer types that account for more than 60 percent of cancer deaths in the U.S. Five of the cancers covered by the test currently have no screening test.
“The use of a combination of selected biomarkers for early detection has the potential to change the way we screen for cancer, and it is based on the same rationale for using combinations of drugs to treat cancers,” says Nickolas Papadopoulos, Ph.D., senior author and professor of oncology and pathology.
The findings were published online by Science on Jan. 18, 2018.
“Circulating tumor DNA mutations can be highly specific markers for cancer. To capitalize on this inherent specificity, we sought to develop a small yet robust panel that could detect at least one mutation in the vast majority of cancers,” says Joshua Cohen, an M.D.-Ph.D. student at the Johns Hopkins University School of Medicine and the paper’s first author. “In fact, keeping the mutation panel small is essential to minimize false-positive results and keep such screening tests affordable.”
The investigators initially explored several hundred genes and 40 protein markers, whittling the number down to segments of 16 genes and eight proteins. They point out that this molecular test is solely aimed at cancer screening and, therefore, is different from other molecular tests, which rely on analyzing large numbers of cancer-driving genes to identify therapeutically actionable targets.
In this study, the test had greater than 99 percent specificity for cancer. “Very high specificity was essential because false-positive results can subject patients to unnecessary invasive follow-up tests and procedures to confirm the presence of cancer,” says Kenneth Kinzler, Ph.D., professor of oncology and co-director of the Ludwig Center. The test was used on 812 healthy controls and produced only seven false-positive results.
The test was evaluated on 1,005 patients with nonmetastatic, stages I to III cancers of the ovary, liver, stomach, pancreas, esophagus, colorectum, lung or breast. The median overall sensitivity, or the ability to find cancer, was 70 percent and ranged from a high of 98 percent for ovarian cancer to a low of 33 percent for breast cancer. For the five cancers that have no screening tests—ovarian, liver, stomach, pancreatic and esophageal cancers—sensitivity ranged from 69 percent to 98 percent.
“A novelty of our classification method is that it combines the probability of observing various DNA mutations together with the levels of several proteins in order to make the final call,” says Cristian Tomasetti, Ph.D., associate professor of oncology and biostatistics, who developed the algorithm. “Another new aspect of our approach is that it uses machine learning to enable the test to accurately determine the location of a tumor down to a small number of anatomic sites in 83 percent of patients.”
Although the current test does not pick up every cancer, it identifies many cancers that would likely otherwise go undetected. “Many of the most promising cancer treatments we have today only benefit a small minority of cancer patients, and we consider them major breakthroughs. If we are going to make progress in early cancer detection, we have to begin looking at it in a more realistic way, recognizing that no test will detect all cancers,” says Bert Vogelstein, M.D., co-director of the Ludwig Center, Clayton Professor of Oncology and Howard Hughes Medical Institute investigator.
To zero in on the analytes they included in their CancerSEEK test, the research team pulled data from more than three decades of cancer genetics research generated at their Ludwig Center at Johns Hopkins, where the first genetic blueprints for cancer were created, as well as data from many other institutions.
To precisely determine the optimal number of DNA bases to assess in the CancerSEEK test, the researchers used a method based on diminishing returns. “The more DNA bases you assay, the more mutations you are capable of finding, but eventually you reach a point of diminishing returns,” explains Cohen. “We designed our test to reflect this point of diminishing returns, including the DNA markers that were useful to detecting the cancers and eliminating those that did not add benefit.” The result was a relatively small panel of highly selective DNA markers.
“This test represents the next step in changing the focus of cancer research from late-stage disease to early disease, which I believe will be critical to reducing cancer deaths in the long term,” says Vogelstein.
CancerSEEK is noninvasive and can, in principle, be administered by primary care providers at the time of other routine blood work. “This has the potential to substantially impact patients. Earlier detection provides many ways to improve outcomes for patients. Optimally, cancers would be detected early enough that they could be cured by surgery alone, but even cancers that are not curable by surgery alone will respond better to systemic therapies when there is less advanced disease,” says Anne Marie Lennon, M.D., Ph.D., associate professor of medicine, surgery and radiology, clinical director of gastroenterology and director of the Multidisciplinary Pancreatic Cyst Program.
The investigators feel that a test that will be used routinely for cancer screening must have a cost in line with or less than other currently available screening tests for single cancers, such as colonoscopy. They envision that the CancerSEEK test will eventually cost less than $500.
Larger studies of the test are currently under way.
In addition to Cohen, Papadopoulos, Lennon, Tomasetti, Kinzler and Vogelstein, other participants include Lu Li, Yuxuan Wang, Christopher Thorburn, Bahman Afsari, Ludmila Danilova, Christopher Douville, Ammar Javed, Fay Wong, Austin Mattox, Ralph Hruban, Christopher Wolfgang, Michael Goggins, Marco Dal Molin, Tian-Li Wang, Richard Roden, Alison Klein, Janine Ptak, Lisa Dobbyn, Joy Schaefer, Natalie Silliman, Maria Popoli, Joshua Vogelstein, James Browne, Robert Schoen, Randall Brand, Jeanne Tie, Peter Gibbs, Hui-Li Wong, Aaron Mansfield, Jin Jen, Samir Hanash, Massimo Falconi, Peter Allen, Shibin Zhou, Chetan Bettegowda and Luis Diaz.
The research was supported by the Lustgarten Foundation for Pancreatic Cancer Research, the Virginia and D.K. Ludwig Fund for Cancer Research, The Commonwealth Fund, the John Templeton Foundation, the Clinomics Program, Mayo Clinic Center for Individualized Medicine, the Mayo Clinic Biobank, the Sol Goldman Center for Pancreatic Cancer Research, The Michael Rolfe Pancreatic Cancer Research Foundation, the Benjamin Baker Scholarship, the Gray Foundation, the Early Detection Research Network, Susan Wojcicki and Dennis Troper, the Marcus Foundation, the Conrad N. Hilton Foundation, the Howard Hughes Medical Institute, and National Institutes of Health Grants P50-CA62924, P50-CA102701, CA06973, GM-07309, and U01CA152753.
COI: Nickolas Papadopoulos, Shibin Zhou, Kenneth Kinzler, Luis Diaz and Bert Vogelstein are founders of Personal Genome Diagnostics, Inc. and PapGene, Inc. Vogelstein and Kinzler are on the Scientific Advisory Board of Sysmex-Inostics, and Vogelstein is on the Scientific Advisory Board of Exelixis GP. These companies and others have licensed technologies from Johns Hopkins, and Papadopoulos, Kinzler, Diaz and Vogelstein receive equity or royalties from these licenses. The terms of these arrangements are being managed by The Johns Hopkins University in accordance with its conflict of interest policies.
More information, including a copy of the paper, can be found online at the Science press package at http://www.eurekalert.org/jrnls/sci. You will need your user ID and password to access this information.


