However, finding GSHs with potential for clinical translation has been as difficult as finding a lunar landing site for a spacecraft — which has to be in smooth and approachable territory, not too steep and surrounded by large hills or cliffs, provide good visibility, and enable a safe return. A GSH, similarly, needs to be accessible by genome editing technologies, free of physical obstacles like genes and other functional sequences, and allow high, stable, and safe expression of a “landed” therapeutic gene.
Thus far, only few candidate GSHs have been explored and they all come with certain caveats. Either they are located in genomic regions that are relatively dense with genes, which means that one or several of them could be compromised in their function by a therapeutic gene inserted in their vicinity, or they contain genes with roles in cancer development that could be inadvertently activated. In addition, candidate GSHs have not been analyzed for the presence of regulatory elements that, although not being genes themselves, can regulate the expression of genes from afar, nor whether inserted genes change global gene expression patterns in cells across the entire genome.
Now, a collaboration of researchers at Harvard’s Wyss Institute for Biologically Inspired Engineering, Harvard Medical School (HMS), and the ETH Zurich in Switzerland, has developed a computational approach to identify GSH sites with significantly higher potential for the safe insertion of therapeutic genes and their durable expression across many cell types. For two out of 2,000 predicted GSH sites, the team provided an in-depth validation with adoptive T cell therapies and in vivo gene therapies for skin diseases in mind. By engineering the identified GSH sites to carry a reporter gene in T cells, and a therapeutic gene in skin cells, respectively, they demonstrated safe and long-lasting expression of the newly introduced genes. The study is published in Cell Reports Methods.
“While GSHs could be utilized as universal landing platforms for gene targeting, and thus expedite the clinical development of gene and cell therapies, so far no site of the human genome has been fully validated and all of them are only acceptable for research applications,” said Wyss Core Faculty member George Church, Ph.D., a senior author on the study. “This makes the collaborative approach that we took toward highly-validated GSHs an important step forward. Together with more effective targeted gene integration tools that we develop in the lab, these GSHs could empower a variety of future clinical translation efforts.” Church is a leader of the Wyss Institute’s Synthetic Biology Platform, and also the Robert Winthrop Professor of Genetics at HMS and Professor of Health Sciences and Technology at Harvard University and the Massachusetts Institute of Technology (MIT).
Together with more effective targeted gene integration tools that we develop in the lab, these GSHs could empower a variety of future clinical translation efforts.
GEORGE CHURCH
Sifting the genome for GSHs
The researchers first set up a computational pipeline that allowed them to predict regions in the genome with potential for use as GSHs by harnessing the wealth of available sequencing data from human cell lines and tissues. “In this step-by-step whole-genome scan we computationally excluded regions encoding proteins, including proteins that have been involved in the formation of tumors, and regions encoding certain types of RNAs with functions in gene expression and other cellular processes. We also eliminated regions that contain so-called enhancer elements, which activate the expression of genes, often from afar, and regions that comprise the centers and ends of chromosomes to avoid mistakes in the replication and segregation of chromosomes during cell division,” said first-author Erik Aznauryan, Ph.D. “This left us with around 2,000 candidate loci all to be further investigated for clinical and biotechnological purposes.”Aznauryan started the project as a graduate student with other members of Sai Reddy’s lab at ETH Zurich’s Department of Biosystems Science and Engineering before he visited the Church lab as part of his graduate work, where he teamed up with Wyss Technology Development Fellow Denitsa Milanova, Ph.D. He since has joined Church’s group as a Postdoctoral Fellow. Reddy, senior and lead author of the collaborative study, is an Associate Professor of Systems and Synthetic Immunology at ETH Zurich and focuses on developing new methods in systems and synthetic biology to engineer immune cells for diverse research and clinical applications.
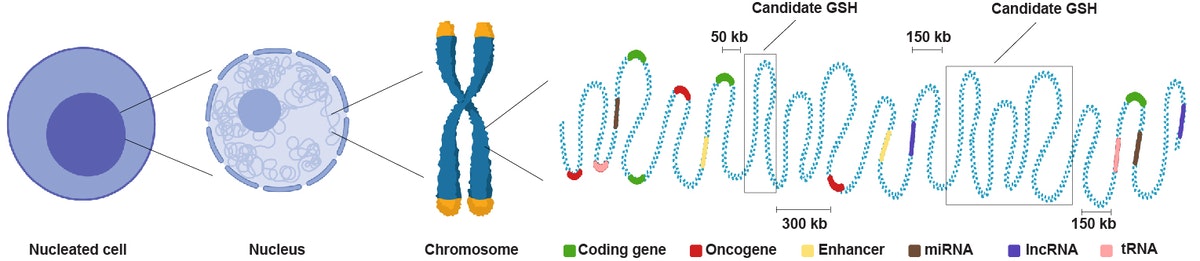
Landing therapeutic genes safely in the human genome
Taking a multi-tiered computational approach, the team scanned the entire human genome landscape for regions that were free of genes encoding proteins, including potentially cancer-promoting proteins, and different types of RNAs, as well as regulatory elements, and that were distanced away from the centers and ends of chromosomes. Selected with these exclusion criteria, the integration of therapeutic genes in candidate GSHs is far less likely to inadvertently inactivate or activate other genes, or cause mistakes in the replication and segregation of chromosomes during cell division. Credit: Wyss Institute at Harvard University
Out of the 2,000 identified GSH sites, the team randomly selected five and investigated them in common human cell lines by inserting reporter genes into each of them using a rapid and efficient CRISPR-Cas9-based genome editing strategy. “Two of the GSH sites allowed particularly high expression of the inserted reporter gene — in fact, significantly higher than expression levels achieved by the team with the same reporter gene engineered into two earlier-generation GSHs. Importantly, the reporter genes harbored by the two GSH sites did not upregulate any cancer-related genes,” said Aznauryan. This also can become possible because regions in the genome distant from one another in the linear DNA sequence of chromosomes, but near in the three-dimensional genome, in which different regions of folded chromosomes touch each other, can become jointly affected when an additional gene is inserted.
Eying clinical translation
To evaluate the two most compelling GSH sites in human cell types with interest for cell and gene therapies, the team investigated them in immune T cells and skin cells, respectively. T cells are used in a number of adoptive cell therapies for the treatment of cancer and autoimmune diseases that could be safer if the receptor-encoding gene was stably inserted into a GSH. Also, skin diseases caused by harmful mutations in genes controlling the function of cells in different skin layers could potentially be cured by insertion and long-term expression of a healthy copy of the mutated gene into a GSH of dividing skin cells that replenish those layers.“We introduced a fluorescent reporter gene into two new GSHs in primary human T cells obtained from blood, and a fully functional LAMB3 gene, an extracellular protein in the skin, into the same GSHs in primary human dermal fibroblasts, and observed long-lasting activity,” said Milanova. “While these GSHs are uniquely positioned to improve on levels and persistence of gene expression in parent and daughter cells for therapeutics, I am particularly excited about emerging ‘gain-of-function’ cellular enhancements that could augment the normal function of cells and organs. The safety aspect is then of paramount importance.” With an entrepreneurial team at the Wyss, Milanova is developing a platform for genetic rejuvenation and enhancements with a focus on skin rejuvenation.
“An extensive sequencing analysis that we undertook in GSH-engineered primary human T cells clearly demonstrated that the insertion has minimal potential for causing tumor-promoting effects, which always is a main concern when genetically modifying cells for therapeutic use,” said Reddy. “The identification of multiple GSH sites, as we have done here, also supports the potential to build more advanced cellular therapies that use multiple transgenes to program sophisticated cellular responses, this is especially relevant in T cell engineering for cancer immunotherapy.”
The identification of multiple GSH sites, as we have done here, also supports the potential to build more advanced cellular therapies that use multiple transgenes to program sophisticated cellular responses, this is especially relevant in T cell engineering for cancer immunotherapy.”“This collaborative interdisciplinary effort demonstrates the power of integrating computational approaches with genome engineering while maintaining a focus on clinical translation. The identification of GSHs in the human genome will greatly augment future developmental therapeutics efforts focused on the engineering of more effective and safer gene and cellular therapies,” said Wyss Founding Director Donald Ingber, M.D., Ph.D., who is also the Judah Folkman Professor of Vascular Biology at HMS and Boston Children’s Hospital, and Professor of Bioengineering at the Harvard John A. Paulson School of Engineering and Applied Sciences.
SAI REDDY, ASSOCIATE PROFESSOR OF SYSTEMS AND SYNTHETIC IMMUNOLOGY AT ETH ZURICH
Additional authors on the study are Alexander Yermanos, Ph.D, and Edo Kapetanovic, members of Reddy’s group; Anna Devaux at the University of Basel, Switzerland; and, Elvira Kinzina at the McGovern Institute for Brain Research at MIT. The study was supported by ETH Research Grants, the Helmut Horten Stiftung and Aging and Longevity-Related Research Fund at HMS, as well as a Genome Engineer Innovation Grant 2019 from Synthego to Aznauryan.
By Benjamin Boettner
wyss.harvard.edu


